BioLayout Express3D is a network visualisation and analysis tool that has been used in the analysis of the FANTOM5 data. It works to identify patterns in data by building network visualisations of correlation matrices i.e. co-expression graphs. Data is loaded into the tool (in a .expression format) which then performs a pairwise all vs. all comparison using the Pearson correlation algorithm. The data may be examined in either direction, by comparing overall the expression patterns between samples or by the individual expression patterns of genes or promoters. Nodes in a graph can therefore either represent samples or genes/promoters, respectively.
Showcased on these pages are a number of graph-based views of the Fantom5 data which are available as webstart files i.e. click and go, or for download and exploration on an installed version of the software. Given the size of the mouse and human data sets (>150k promoters, 100’s of samples) we illustrate only subsections of the Fantom5 data due to the potential size of the graphs and the issues with rendering and interpreting such large networks. This is possible using BioLayout Express3D but requires a higher than normal specification of hardware.
To better understand how to navigate and explore these graphs please click here.
Human Primary Cells – Promoters for transcription factors only
These data represent all CAGE data mapping to the promoters of all known human transcription factors as analysed in the primary cell samples. In total this represents 7,596 promoters from 1,606 transcription factors whose expression has been mapped across 507 samples of primary cells. The graph has been constructed using a Pearson correlation value of r = 0.7 which connects 4,556 promoters (57,630 edges) by virtue of their coexpression at this relatively conservative correlation value and clustered using an MCL inflation value of 2.2.
This graph provides a resource for the exploration of the transcription factors that drive differentiation in human cells
Human Primary Cells - A sample-to-sample correlation matrix
These data represent a comparison of all CAGE data from the 507 human primary cell samples when compared to each other. A sample-to-sample correlation matrix was calculated comparing the data derived from each sample to the others. This matrix was then filtered to included only those sample-to-sample relationships greater than r = 0.8. Of the 507 samples 497 are included in the graph at this threshold connected by 6,284 edges and the graph clustered using an MCL inflation value of 2.2. 2D and 3D layouts of the graphs are provided.
These graphs provide a resource for the exploration of overall similarity between transcriptional landscape of human cells.
Mouse Cell and Tissues - Promoters for transcription factors only
These data represent all CAGE data mapping to the promoters of all known mouse transcription factors as analysed in the primary cell samples. In total this represents 4,906 promoters from 1,362 transcription factors whose expression has been mapped across 358 samples of mouse cells and tissue. The graph has been constructed using a Pearson correlation value of r = 0.7 which connects 3,768 promoters (79,409 edges) by virtue of their coexpression at this relatively conservative correlation value and clustered using an MCL inflation value of 2.2.
This graph provides a resource for the exploration of the transcription factors that drive differentiation in mouse tissues and cells.
Mouse Cell and Tissues - A sample-to-sample correlation matrix
These data represent a comparison of all CAGE data from the 358 mouse cell and tissue samples when compared to each other. A sample-to-sample correlation matrix was calculated comparing the data derived from each sample to the others. This matrix was then filtered to included only those sample-to-sample relationships greater than r = 0.8. Of the 358 samples 331 are included in the graph at this threshold connected by 1,815 edges and the graph clustered using an MCL inflation value of 2.2.
These graphs provide a resource for the exploration of overall similarity between transcriptional landscape of mouse cells and tissues.
Coexpression clustering of all human promoters in FANTOM5
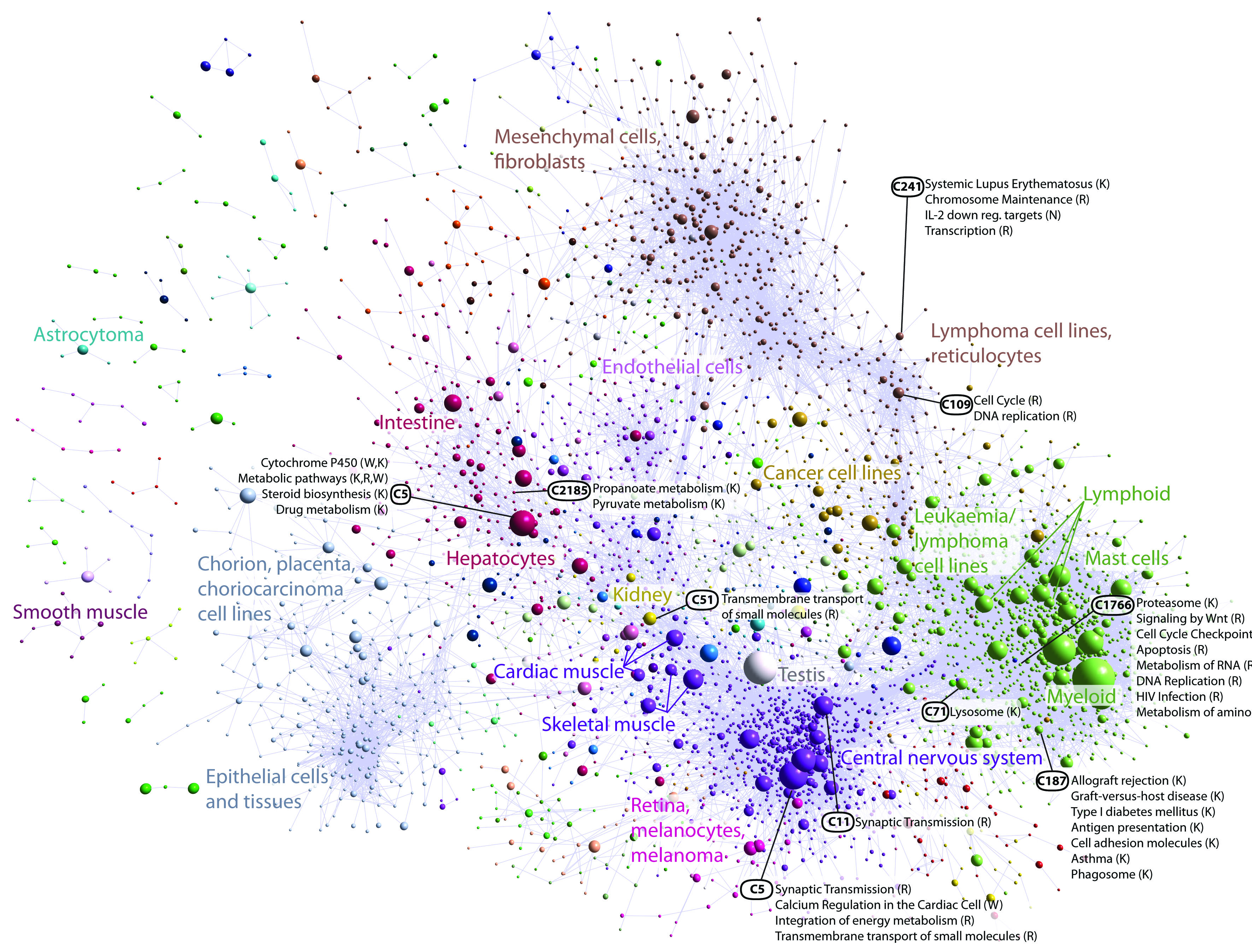
Graph associated with Figure 4 from promoterome paper: Coexpression clustering of human promoters in FANTOM5. Collapsed coexpression network derived from 4,882 coexpression groups (one node is one group of promoters; 4,664 groups are shown here) derived from expression profiles of 124,090 promoters across all primary cell types, tissues and cell lines (visualized using Biolayout Express3D, r.0.75, MCLi 2.2). For display, each group of promoters is collapsed into a sphere, the radius of which is proportional to the cube root of the number of promoters in that group. Edges indicate r.0.6 between the average expression profiles of each cluster. Colours indicate loosely-associated collections of coexpression groups (MCLi1.2). Labels show representative descriptions of the dominant cell type in coexpression groups in each region of the network. Promoters and genes in the coexpression groups are available online at (https://fantom.gsc.riken.jp/5/data/).
Direct link to the data directory:
